11 Байесовский регрессионный анализ
library(tidyverse)11.1 Основы регрессионного анализа
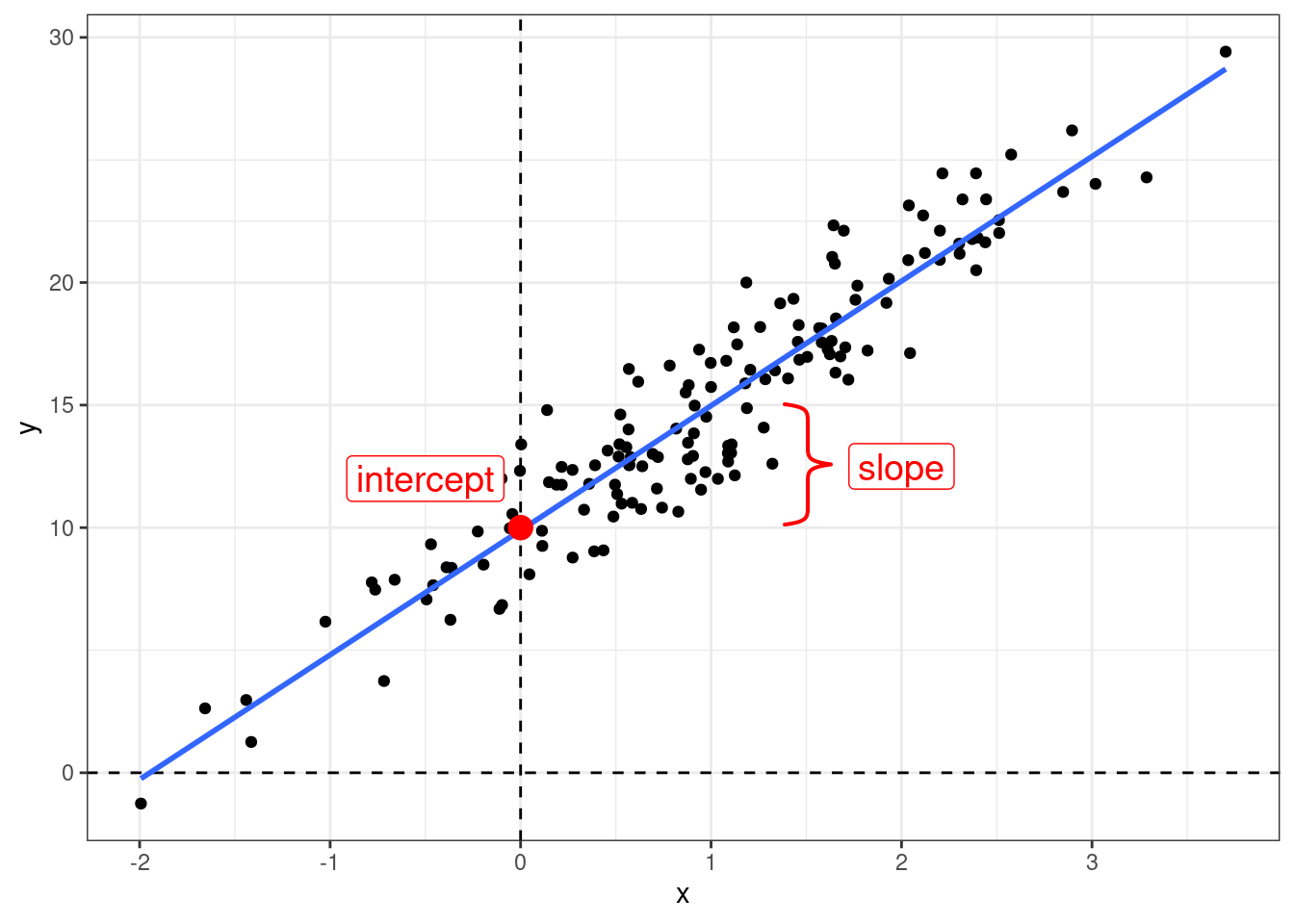
Когда мы используем регрессионный анализ, мы пытаемся оценить два параметра:
- свободный член (intercept) – значение \(y\) при \(x = 0\);
- угловой коэффициент (slope) – изменение \(y\) при изменении \(x\) на одну единицу.
\[y_i = \beta_0 + \beta_1\times x_i + \epsilon_i\]
Причем, иногда мы можем один или другой параметр считать равным нулю.
При этом, вне зависимости от статистической школы, у регрессии есть свои ограничения на применение:
- линейность связи между \(x\) и \(y\);
- нормальность распределение остатков \(\epsilon_i\);
- гомоскидастичность — равномерность распределения остатков на всем протяжении \(x\);
- независимость переменных;
- независимость наблюдений друг от друга.
11.2 brms
Для анализа возьмем датасет, который я составил из UD-корпусов и попробуем смоделировать связь между количеством слов в тексте и количеством уникальных слов (закон Хердана-Хипса).
ud <- read_csv("https://raw.githubusercontent.com/agricolamz/udpipe_count_n_words_and_tokens/master/filtered_dataset.csv")
glimpse(ud)Rows: 20,705
Columns: 5
$ doc_id <chr> "KR1d0052_001", "KR1d0052_002", "KR1d0052_003", "KR1d0052_…
$ n_words <dbl> 3516, 2131, 4927, 4884, 4245, 5027, 3406, 2202, 2673, 2300…
$ n_tokens <dbl> 842, 546, 869, 883, 737, 1085, 494, 443, 573, 578, 660, 87…
$ language <chr> "Classical_Chinese", "Classical_Chinese", "Classical_Chine…
$ corpus_code <chr> "Kyoto", "Kyoto", "Kyoto", "Kyoto", "Kyoto", "Kyoto", "Kyo…Для начала, нарушим кучу ограничений на применение регрессии и смоделируем модель для вот таких вот данных, взяв только тексты меньше 1500 слов:
ud %>%
filter(n_words < 1500) ->
ud
ud %>%
ggplot(aes(n_words, n_tokens))+
geom_point()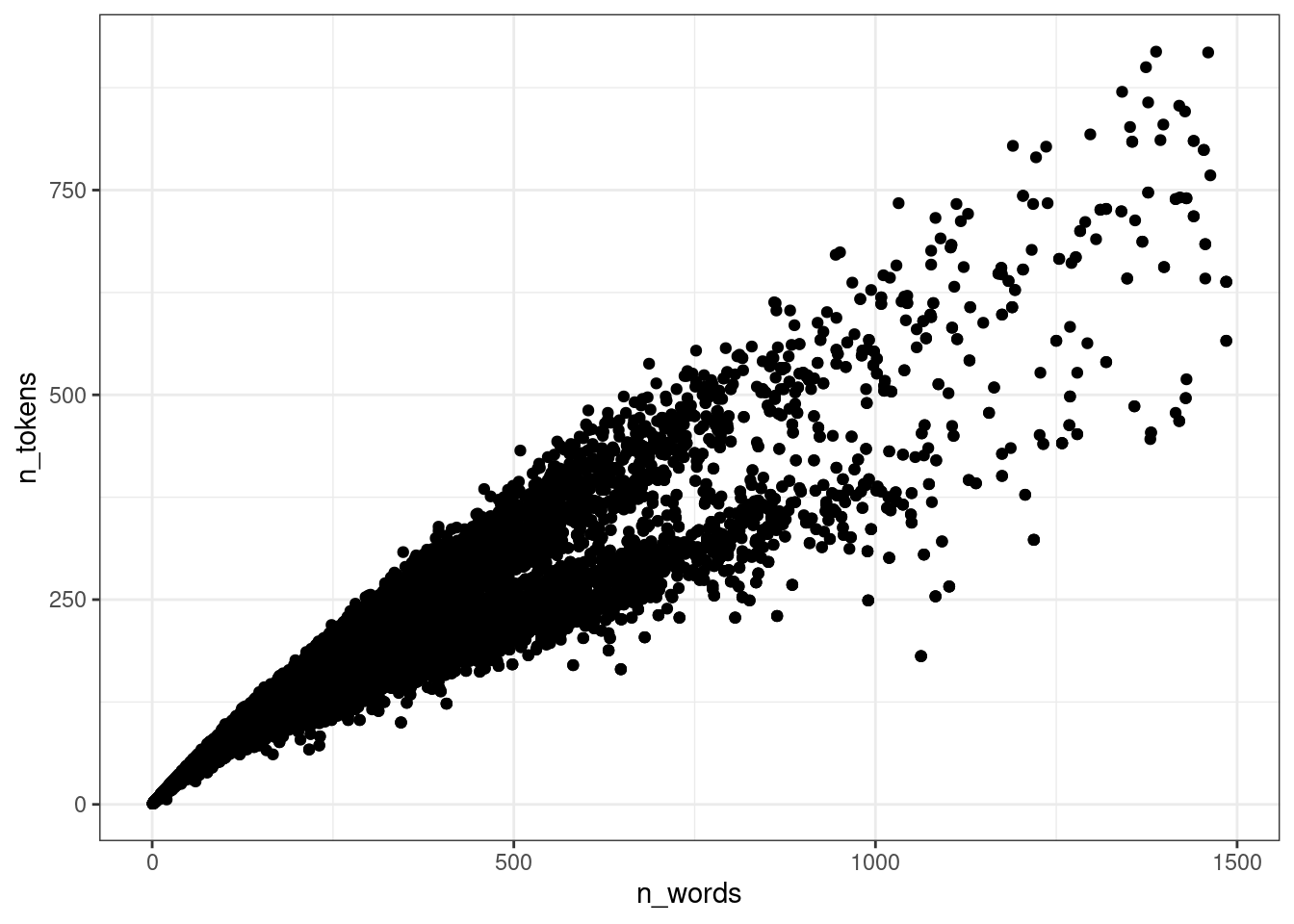
11.2.1 Модель только со свободным членом
library(brms)
parallel::detectCores()[1] 8n_cores <- 7 # parallel::detectCores() - 1
get_prior(n_tokens ~ 1,
data = ud)Вот модель с встроенными априорными распределениями:
fit_intercept <- brm(n_tokens ~ 1,
data = ud,
cores = n_cores, refresh = 0, silent = TRUE)Running /usr/lib/R/bin/R CMD SHLIB foo.c
gcc -std=gnu99 -std=gnu11 -I"/usr/share/R/include" -DNDEBUG -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/Rcpp/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/unsupported" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/BH/include" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/src/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppParallel/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/rstan/include" -DEIGEN_NO_DEBUG -DBOOST_DISABLE_ASSERTS -DBOOST_PENDING_INTEGER_LOG2_HPP -DSTAN_THREADS -DBOOST_NO_AUTO_PTR -include '/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp' -D_REENTRANT -DRCPP_PARALLEL_USE_TBB=1 -fpic -g -O2 -fdebug-prefix-map=/build/r-base-i2PIHO/r-base-4.1.2=. -fstack-protector-strong -Wformat -Werror=format-security -Wdate-time -D_FORTIFY_SOURCE=2 -g -c foo.c -o foo.o
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:88,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:1: error: unknown type name ‘namespace’
628 | namespace Eigen {
| ^~~~~~~~~
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:17: error: expected ‘=’, ‘,’, ‘;’, ‘asm’ or ‘__attribute__’ before ‘{’ token
628 | namespace Eigen {
| ^
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:96:10: fatal error: complex: No such file or directory
96 | #include <complex>
| ^~~~~~~~~
compilation terminated.
make: *** [/usr/lib/R/etc/Makeconf:168: foo.o] Error 1При желании встроенные априорные расспеределения можно не использовать и вставлять в аргумент prior априорные распределения по вашему желанию.
fit_intercept Family: gaussian
Links: mu = identity; sigma = identity
Formula: n_tokens ~ 1
Data: ud (Number of observations: 20282)
Draws: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup draws = 4000
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 135.65 0.85 134.00 137.30 1.00 2734 2548
Family Specific Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sigma 121.75 0.60 120.58 122.93 1.00 3767 2604
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).plot(fit_intercept)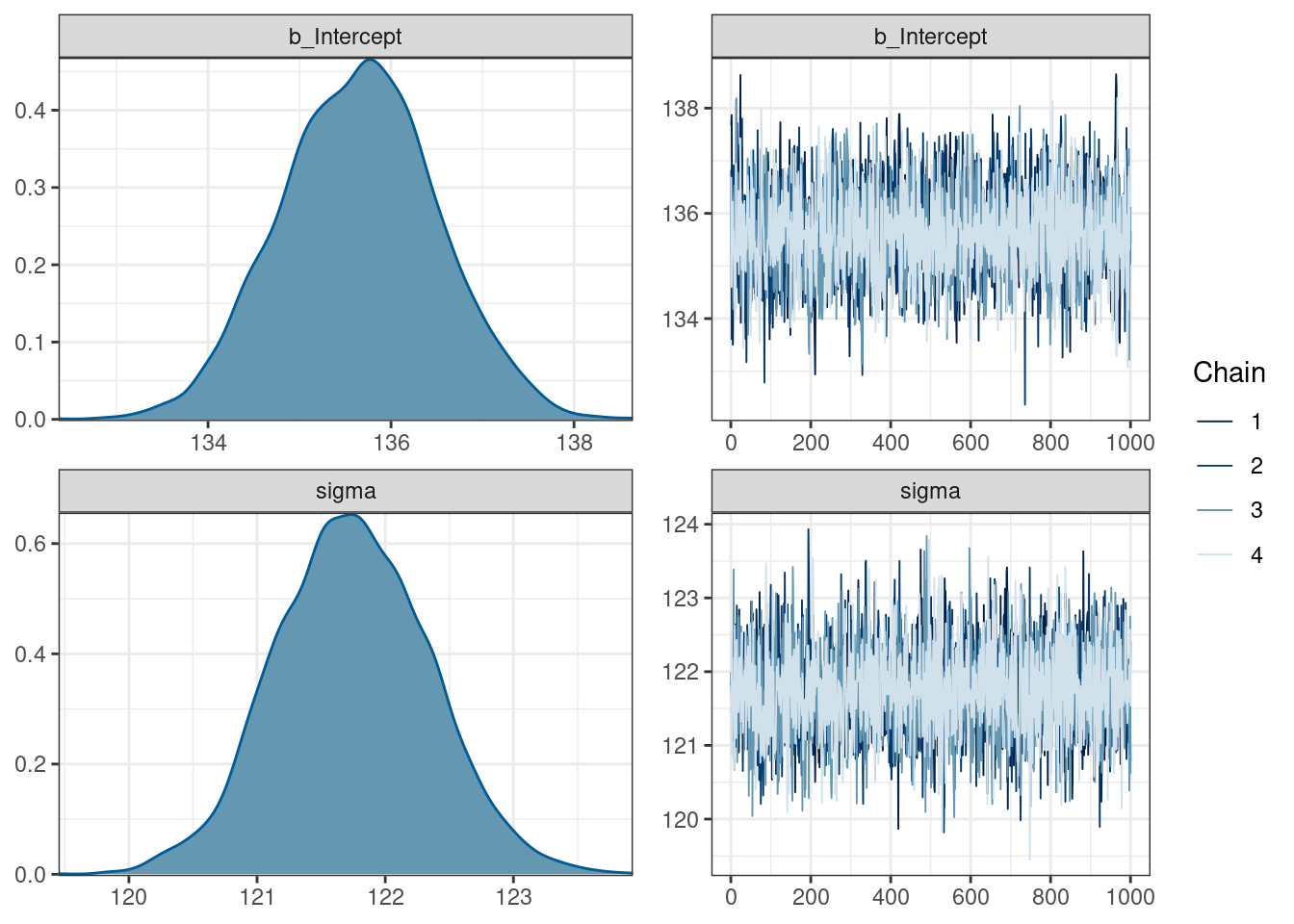
Взято остюда:
11.2.1.1 R-hat
R-hat convergence diagnostic compares the between- and within-chain estimates for model parameters and other univariate quantities of interest. If chains have not mixed well (ie, the between- and within-chain estimates don’t agree), R-hat is larger than 1. We recommend running at least four chains by default and only using the sample if R-hat is less than 1.01. Stan reports R-hat which is the maximum of rank normalized split-R-hat and rank normalized folded-split-R-hat, which works for thick tailed distributions and is sensitive also to differences in scale. For more details on this diagnostic, see https://arxiv.org/abs/1903.08008.
Recommendations:
- Look at Bulk- and Tail-ESS for further information.
- Look at the rank plot to see how the chains differ from each other.
- Look at the local and quantile efficiency plots.
- You might try setting a higher value for the iter argument. By default iter is 2000
11.2.1.2 Bulk ESS
Roughly speaking, the effective sample size (ESS) of a quantity of interest captures how many independent draws contain the same amount of information as the dependent sample obtained by the MCMC algorithm. Clearly, the higher the ESS the better. Stan uses R-hat adjustment to use the between-chain information in computing the ESS. For example, in case of multimodal distributions with well-separated modes, this leads to an ESS estimate that is close to the number of distinct modes that are found.
Bulk-ESS refers to the effective sample size based on the rank normalized draws. This does not directly compute the ESS relevant for computing the mean of the parameter, but instead computes a quantity that is well defined even if the chains do not have finite mean or variance. Overall bulk-ESS estimates the sampling efficiency for the location of the distribution (e.g. mean and median).
Often quite smaller ESS would be sufficient for the desired estimation accuracy, but the estimation of ESS and convergence diagnostics themselves require higher ESS. We recommend requiring that the bulk-ESS is greater than 100 times the number of chains. For example, when running four chains, this corresponds to having a rank-normalized effective sample size of at least 400.
Recommendations:
- You might try setting a higher value for the iter argument. By default iter is 2000
- Look at the rank plot to see how the chains differ from each other.
- Look at the local and quantile efficiency plots.
- Look at change in bulk-ESS when the number of iterations increase. If R-hat is less than 1.01 and bulk-ESS grows linearly with the number of iterations and eventually exceeds the recommended limit, the mixing is sufficient but MCMC has high autocorrelation requiring a large number of iterations
11.2.1.3 Tail ESS
Tail-ESS computes the minimum of the effective sample sizes (ESS) of the 5% and 95% quantiles. Tail-ESS can help diagnosing problems due to different scales of the chains and slow mixing in the tails. See also general information about ESS above in description of bulk-ESS.
Recommendations:
- You might try setting a higher value for the iter argument. By default iter is 2000
- Look at the rank plot to see how the chains differ from each other.
- Look at the local and quantile efficiency plots.
- Look at change in tail-ESS when the number of iterations increase. If R-hat is less than 1.01 and tail-ESS grows linearly with the number of iterations and eventually exceeds the recommended limit, the mixing is sufficient but MCMC has high autocorrelation requiring a large number of iterations
11.2.2 Модель только с угловым коэффициентом
get_prior(n_tokens ~ n_words+0,
data = ud)fit_slope <- brm(n_tokens ~ n_words+0,
data = ud,
cores = n_cores, refresh = 0, silent = TRUE)Running /usr/lib/R/bin/R CMD SHLIB foo.c
gcc -std=gnu99 -std=gnu11 -I"/usr/share/R/include" -DNDEBUG -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/Rcpp/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/unsupported" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/BH/include" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/src/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppParallel/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/rstan/include" -DEIGEN_NO_DEBUG -DBOOST_DISABLE_ASSERTS -DBOOST_PENDING_INTEGER_LOG2_HPP -DSTAN_THREADS -DBOOST_NO_AUTO_PTR -include '/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp' -D_REENTRANT -DRCPP_PARALLEL_USE_TBB=1 -fpic -g -O2 -fdebug-prefix-map=/build/r-base-i2PIHO/r-base-4.1.2=. -fstack-protector-strong -Wformat -Werror=format-security -Wdate-time -D_FORTIFY_SOURCE=2 -g -c foo.c -o foo.o
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:88,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:1: error: unknown type name ‘namespace’
628 | namespace Eigen {
| ^~~~~~~~~
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:17: error: expected ‘=’, ‘,’, ‘;’, ‘asm’ or ‘__attribute__’ before ‘{’ token
628 | namespace Eigen {
| ^
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:96:10: fatal error: complex: No such file or directory
96 | #include <complex>
| ^~~~~~~~~
compilation terminated.
make: *** [/usr/lib/R/etc/Makeconf:168: foo.o] Error 1fit_slope Family: gaussian
Links: mu = identity; sigma = identity
Formula: n_tokens ~ n_words + 0
Data: ud (Number of observations: 20282)
Draws: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup draws = 4000
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
n_words 0.53 0.00 0.53 0.53 1.00 4360 3010
Family Specific Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sigma 42.79 0.21 42.40 43.20 1.01 1180 1357
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).plot(fit_slope)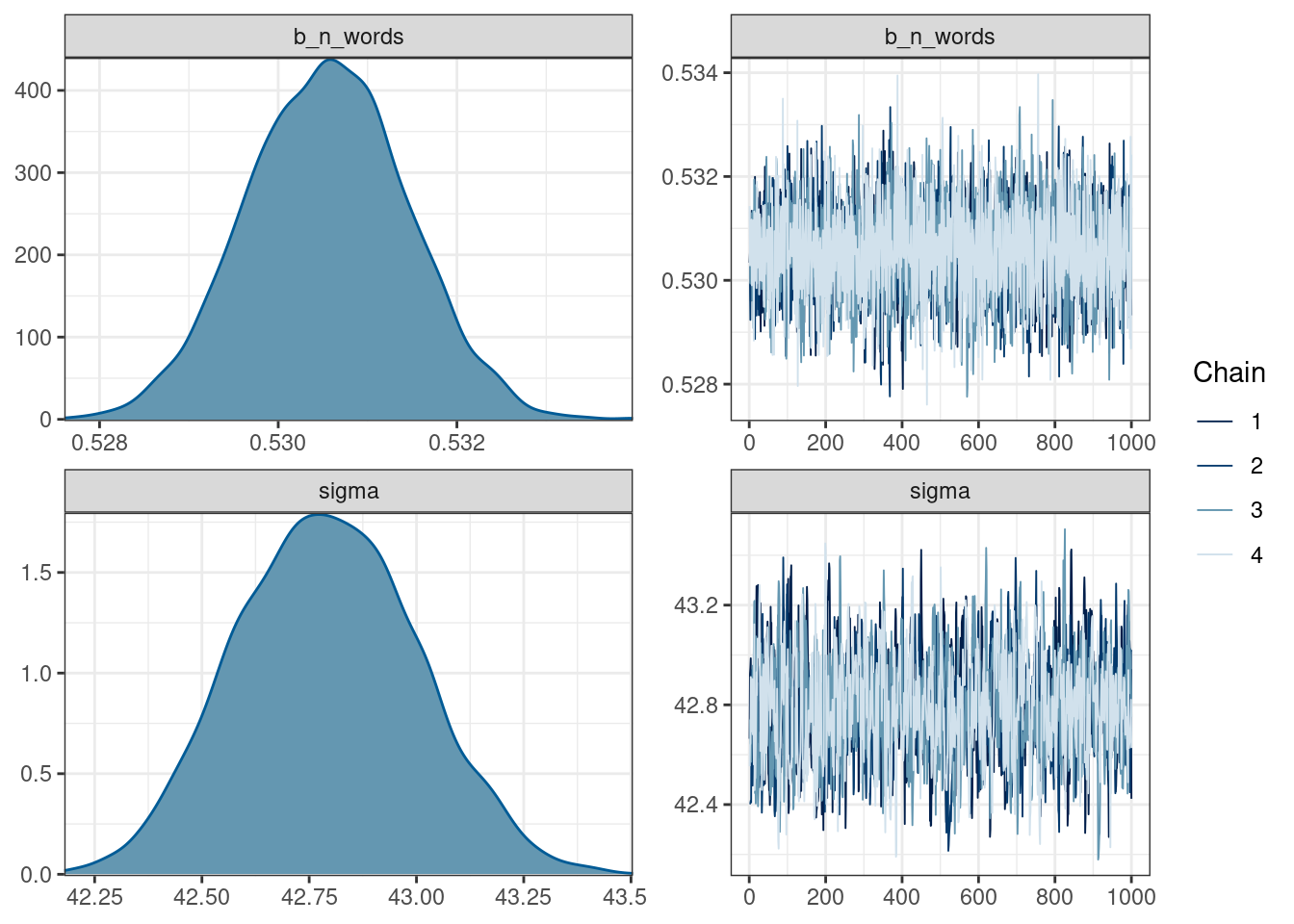
Давайте совместим предсказания модели и наши наблюдения.
library(tidybayes)
ud %>%
add_epred_draws(fit_slope, ndraws = 50) %>%
ggplot(aes(n_words, n_tokens))+
geom_point(alpha = 0.01)+
stat_lineribbon(aes(y = .epred), color = "red") 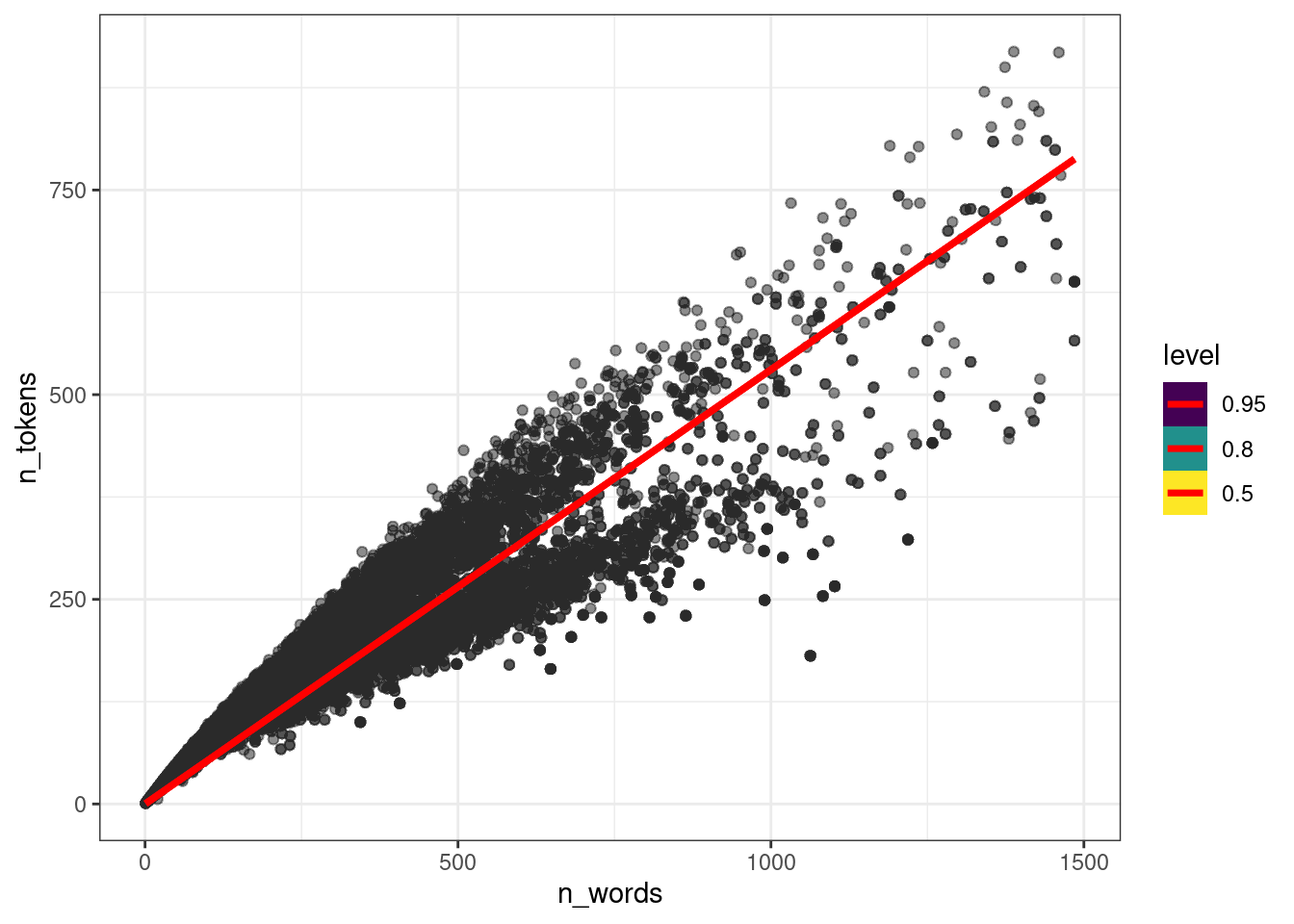
11.2.3 Модель с угловым коэффициентом и свободным членом
get_prior(n_tokens ~ n_words,
data = ud)fit_slope_intercept <- brm(n_tokens ~ n_words,
data = ud,
cores = n_cores, refresh = 0, silent = TRUE)Running /usr/lib/R/bin/R CMD SHLIB foo.c
gcc -std=gnu99 -std=gnu11 -I"/usr/share/R/include" -DNDEBUG -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/Rcpp/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/unsupported" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/BH/include" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/src/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppParallel/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/rstan/include" -DEIGEN_NO_DEBUG -DBOOST_DISABLE_ASSERTS -DBOOST_PENDING_INTEGER_LOG2_HPP -DSTAN_THREADS -DBOOST_NO_AUTO_PTR -include '/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp' -D_REENTRANT -DRCPP_PARALLEL_USE_TBB=1 -fpic -g -O2 -fdebug-prefix-map=/build/r-base-i2PIHO/r-base-4.1.2=. -fstack-protector-strong -Wformat -Werror=format-security -Wdate-time -D_FORTIFY_SOURCE=2 -g -c foo.c -o foo.o
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:88,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:1: error: unknown type name ‘namespace’
628 | namespace Eigen {
| ^~~~~~~~~
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:17: error: expected ‘=’, ‘,’, ‘;’, ‘asm’ or ‘__attribute__’ before ‘{’ token
628 | namespace Eigen {
| ^
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:96:10: fatal error: complex: No such file or directory
96 | #include <complex>
| ^~~~~~~~~
compilation terminated.
make: *** [/usr/lib/R/etc/Makeconf:168: foo.o] Error 1fit_slope_intercept Family: gaussian
Links: mu = identity; sigma = identity
Formula: n_tokens ~ n_words
Data: ud (Number of observations: 20282)
Draws: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup draws = 4000
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 24.31 0.37 23.60 25.03 1.00 2919 2740
n_words 0.48 0.00 0.48 0.48 1.00 4859 2569
Family Specific Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sigma 39.05 0.19 38.67 39.44 1.00 2290 2065
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).plot(fit_slope_intercept)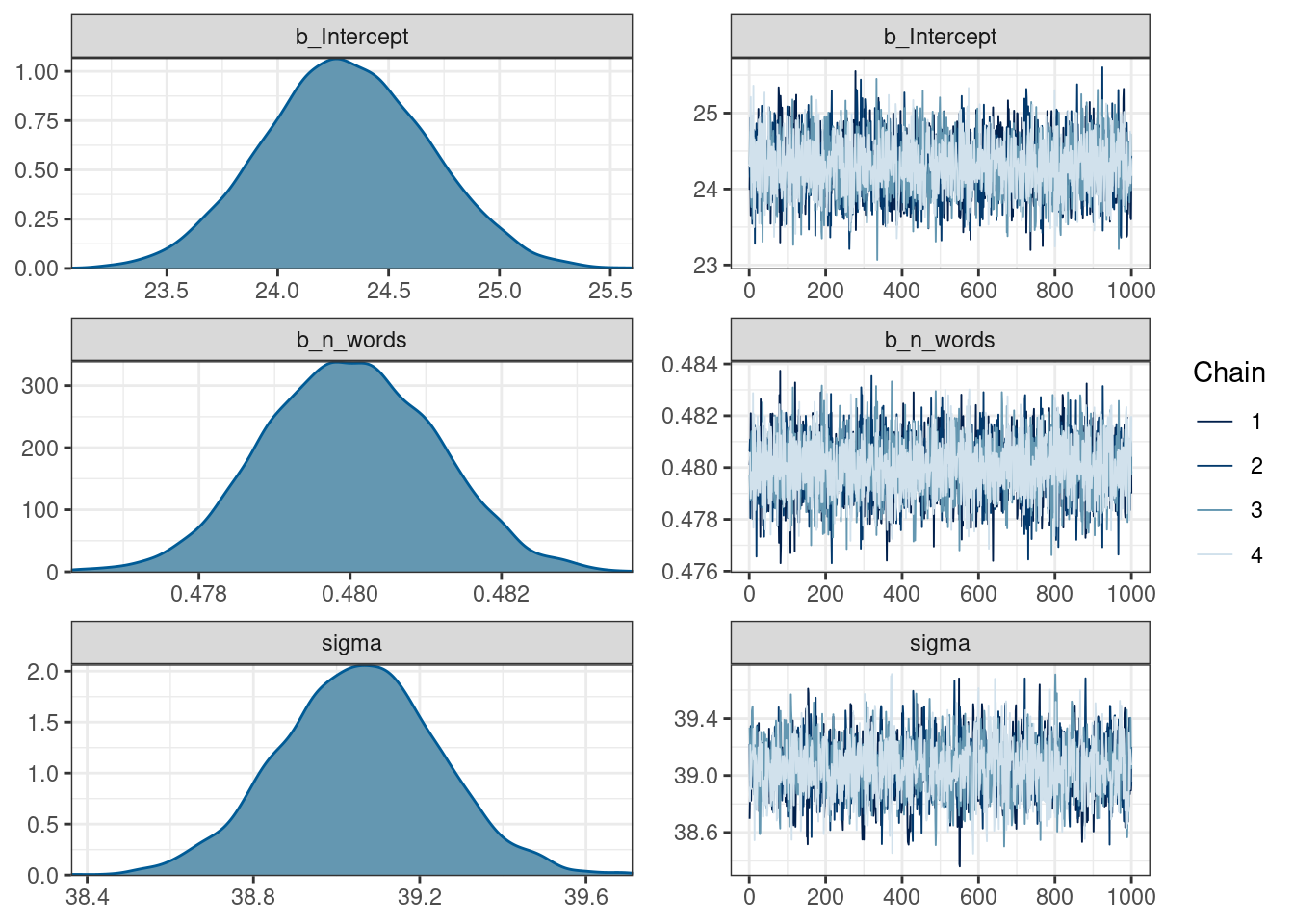
ud %>%
add_epred_draws(fit_slope_intercept, ndraws = 50) %>%
ggplot(aes(n_words, n_tokens))+
geom_point(alpha = 0.01)+
stat_lineribbon(aes(y = .epred), color = "red") 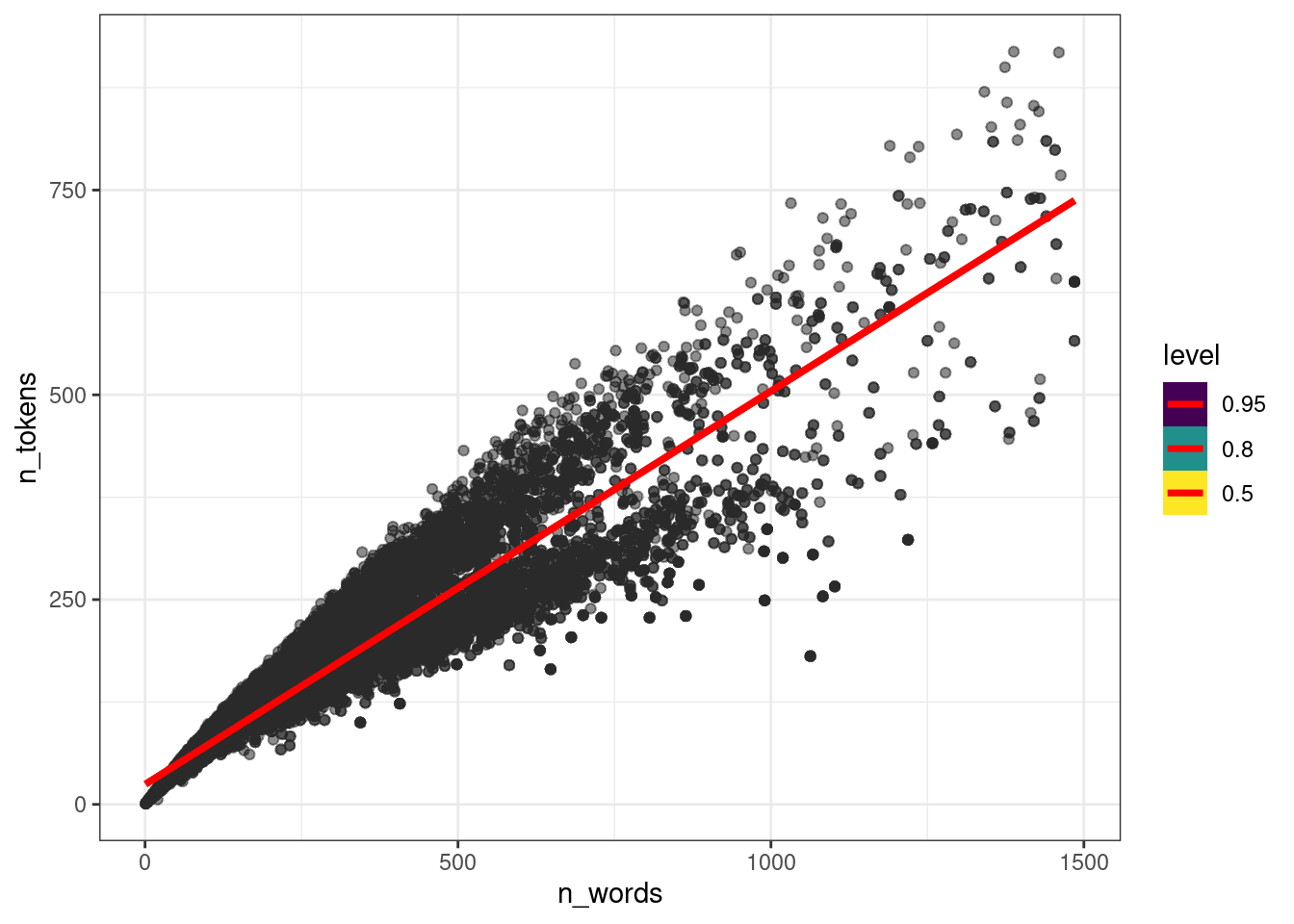
11.2.4 Модель со смешанными эффектами
В данных есть группировка по языкам, которую мы все это время игнорировали. Давайте сделаем модель со смешанными эффектами:
get_prior(n_tokens ~ n_words+(1|language),
data = ud)fit_mixed <- brm(n_tokens ~ n_words + (1|language),
data = ud,
cores = n_cores, refresh = 0, silent = TRUE)Running /usr/lib/R/bin/R CMD SHLIB foo.c
gcc -std=gnu99 -std=gnu11 -I"/usr/share/R/include" -DNDEBUG -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/Rcpp/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/unsupported" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/BH/include" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/src/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppParallel/include/" -I"/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/rstan/include" -DEIGEN_NO_DEBUG -DBOOST_DISABLE_ASSERTS -DBOOST_PENDING_INTEGER_LOG2_HPP -DSTAN_THREADS -DBOOST_NO_AUTO_PTR -include '/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp' -D_REENTRANT -DRCPP_PARALLEL_USE_TBB=1 -fpic -g -O2 -fdebug-prefix-map=/build/r-base-i2PIHO/r-base-4.1.2=. -fstack-protector-strong -Wformat -Werror=format-security -Wdate-time -D_FORTIFY_SOURCE=2 -g -c foo.c -o foo.o
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:88,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:1: error: unknown type name ‘namespace’
628 | namespace Eigen {
| ^~~~~~~~~
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/src/Core/util/Macros.h:628:17: error: expected ‘=’, ‘,’, ‘;’, ‘asm’ or ‘__attribute__’ before ‘{’ token
628 | namespace Eigen {
| ^
In file included from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Dense:1,
from /home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/StanHeaders/include/stan/math/prim/mat/fun/Eigen.hpp:13,
from <command-line>:
/home/agricolamz/R/x86_64-pc-linux-gnu-library/4.1/RcppEigen/include/Eigen/Core:96:10: fatal error: complex: No such file or directory
96 | #include <complex>
| ^~~~~~~~~
compilation terminated.
make: *** [/usr/lib/R/etc/Makeconf:168: foo.o] Error 1fit_mixed Family: gaussian
Links: mu = identity; sigma = identity
Formula: n_tokens ~ n_words + (1 | language)
Data: ud (Number of observations: 20282)
Draws: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup draws = 4000
Group-Level Effects:
~language (Number of levels: 9)
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sd(Intercept) 64.03 18.87 38.13 111.85 1.01 558 1054
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 5.91 20.69 -36.51 46.11 1.00 642 824
n_words 0.49 0.00 0.49 0.49 1.00 4075 2333
Family Specific Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sigma 29.28 0.14 29.01 29.55 1.00 2316 1988
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).plot(fit_mixed)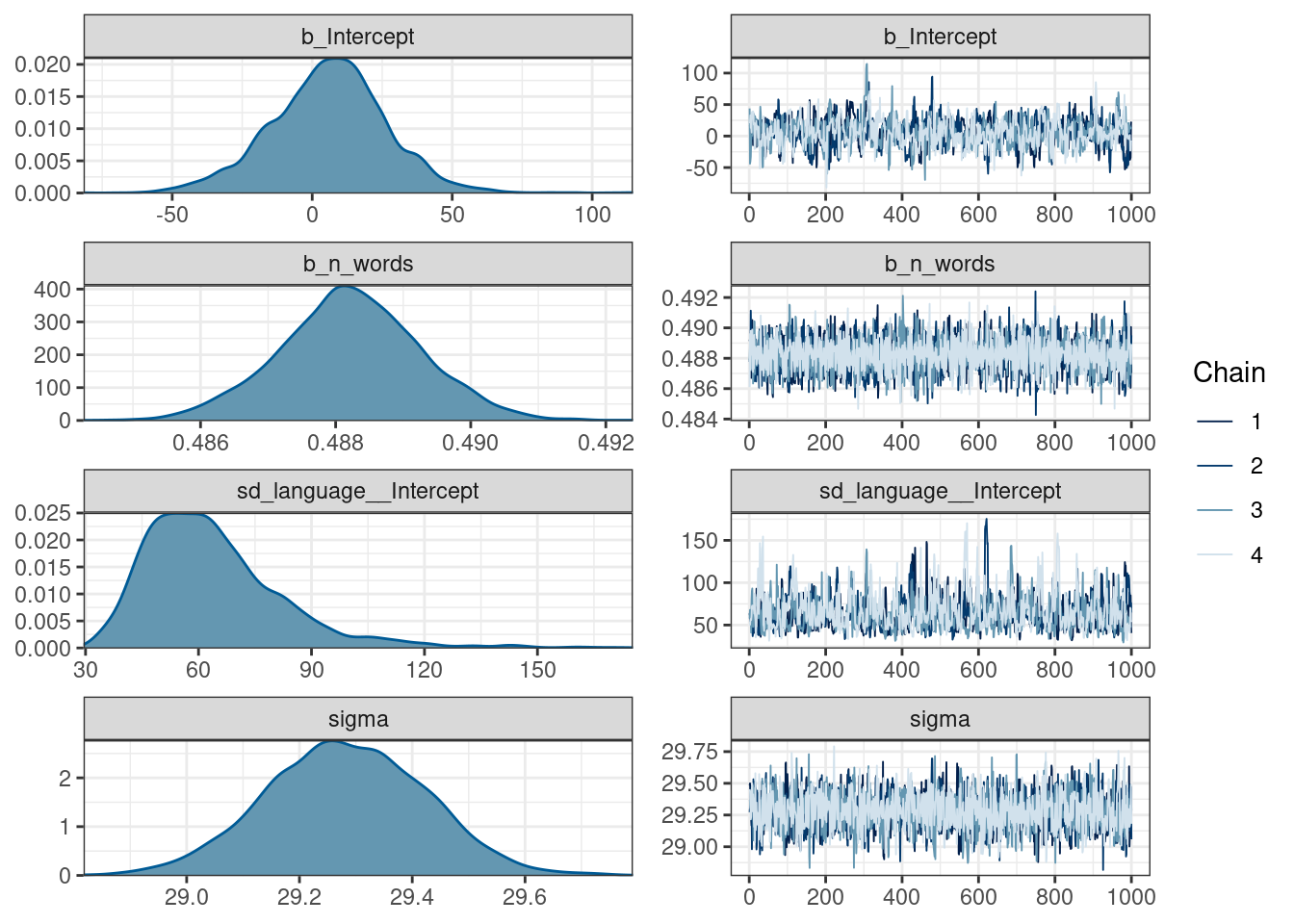
ud %>%
add_epred_draws(fit_mixed, ndraws = 50) %>%
ggplot(aes(n_words, n_tokens))+
geom_point(alpha = 0.01)+
stat_lineribbon(aes(y = .epred), color = "red") +
facet_wrap(~language)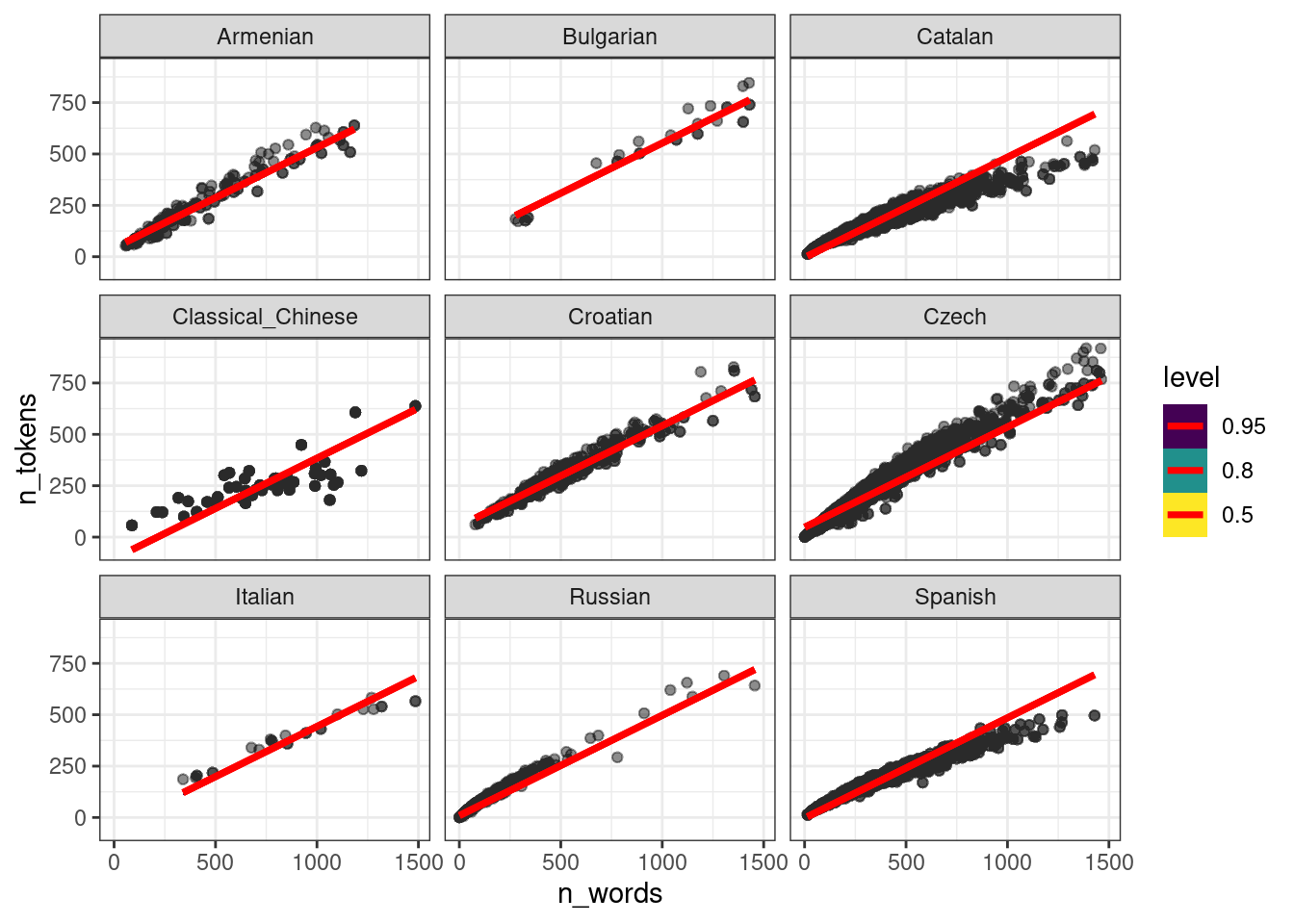
То, что получилось учитывает общий эффект всех языков: посмотрите на каталанский. Если построить модель по каждому языку, то получится совсем другая картина:
ud %>%
ggplot(aes(n_words, n_tokens))+
geom_smooth(method = "lm") +
geom_point(alpha = 0.3)+
facet_wrap(~language)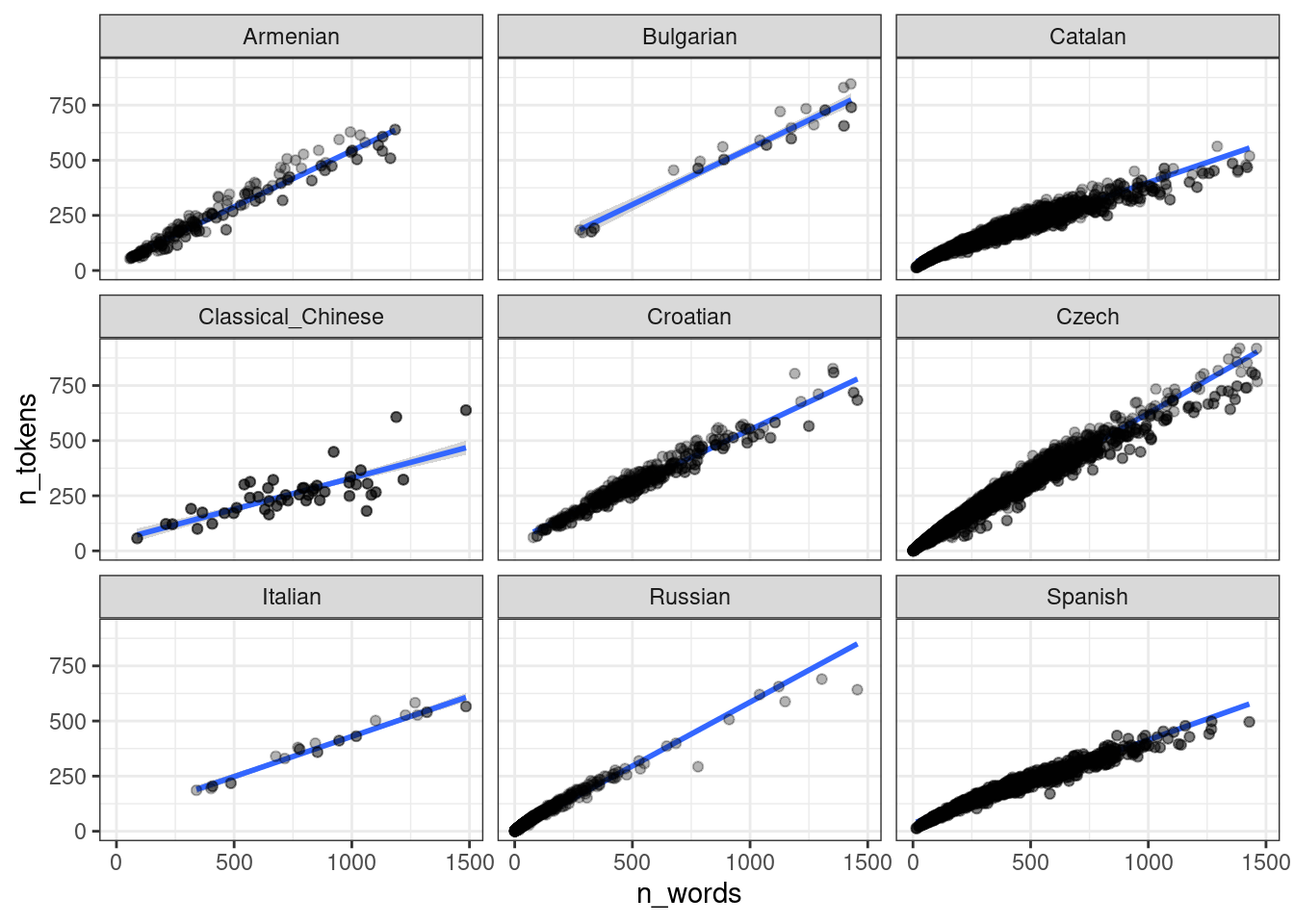
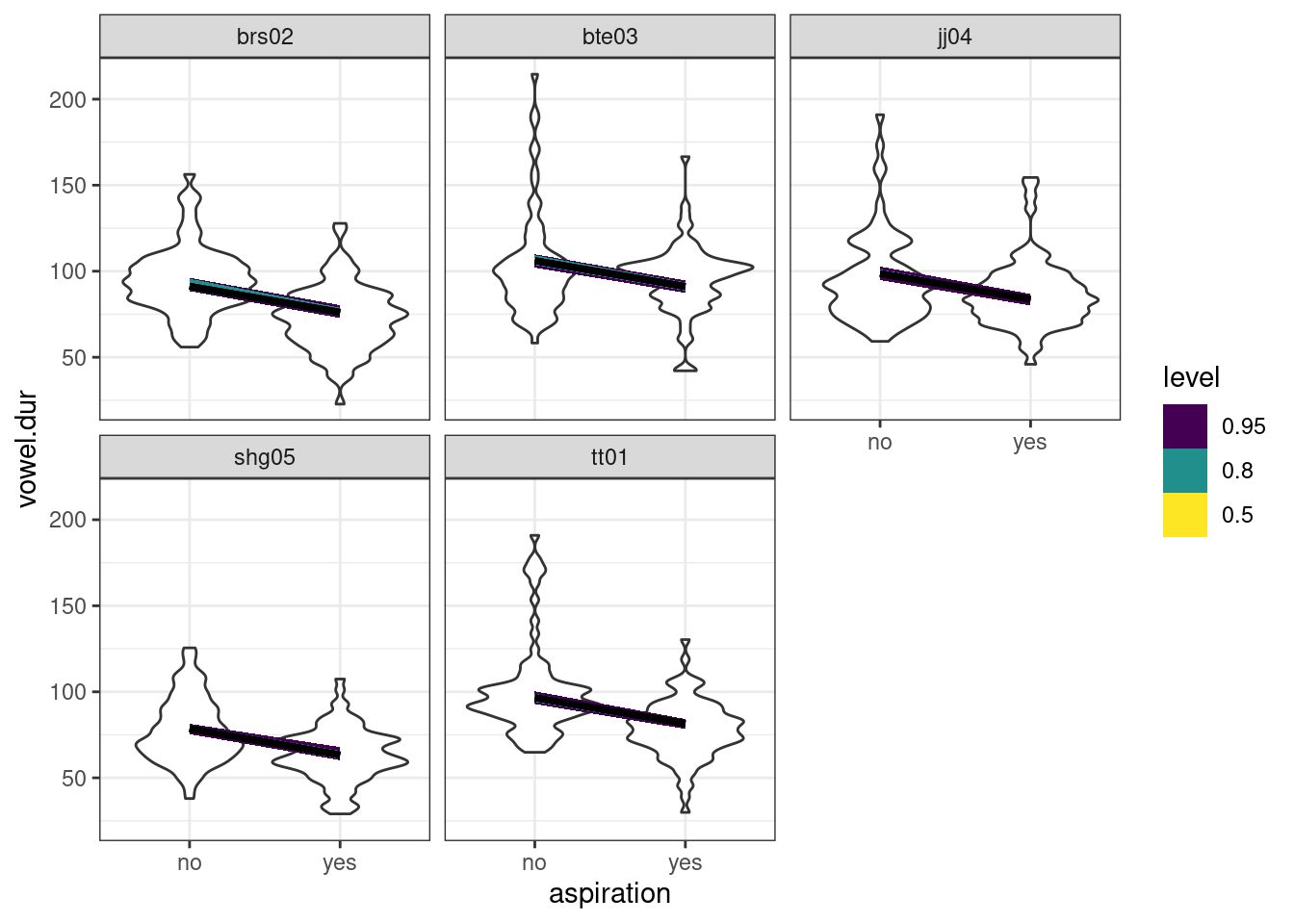
В работе (Coretta 2016) собраны данные длительности исландских гласных. Используя байесовскую регрессию, смоделируйте длительность гласного (vowel.dur) в зависимости от аспирированности (aspiration) учитывая эффект носителя. Визуализируйте результаты модели